MA Plot
The MA Plot app provides an intuitive way to inspect and directly compare log fold change values generated during differential expression analysis. In PanHunter, the outcomes of these analyses are saved as Comparisons or Comparison Groups. For additional details on differential expression analysis or instructions on creating a comparison in PanHunter, please refer to the Algorithms section and the New Comparison app documentation.
The side‑by‑side visualization makes it easy to quickly identify genes of interest. In many cases, genes that exhibit both a high log fold change and a high expression level are particularly noteworthy — especially when they show the same direction of expression change across multiple comparisons.
Have a look at the quick app‑tour video below:
Getting started with MA Plot
The MA Plot interface follows the standard PanHunter layout and consists of the following compontents:
- Tools panel at the left-hand side
- Data selection
- Plot options
Available in Plots and Detailed Plot views - Features to highlight
Available in Plots and Detailed Plot views
- Results view section to the right
- Metadata table
- Plots
- Detailed plot
- General widgets
- Comparison Selector
- Table widget
- Interactive plots
- Plotly plots
Comparison Selection
The MA Plot allows you to visually compare two Comparisons side by side. When you open the app, no data is preselected. Use the Comparison Selection module on the left side to choose comparisons you want to analyze.
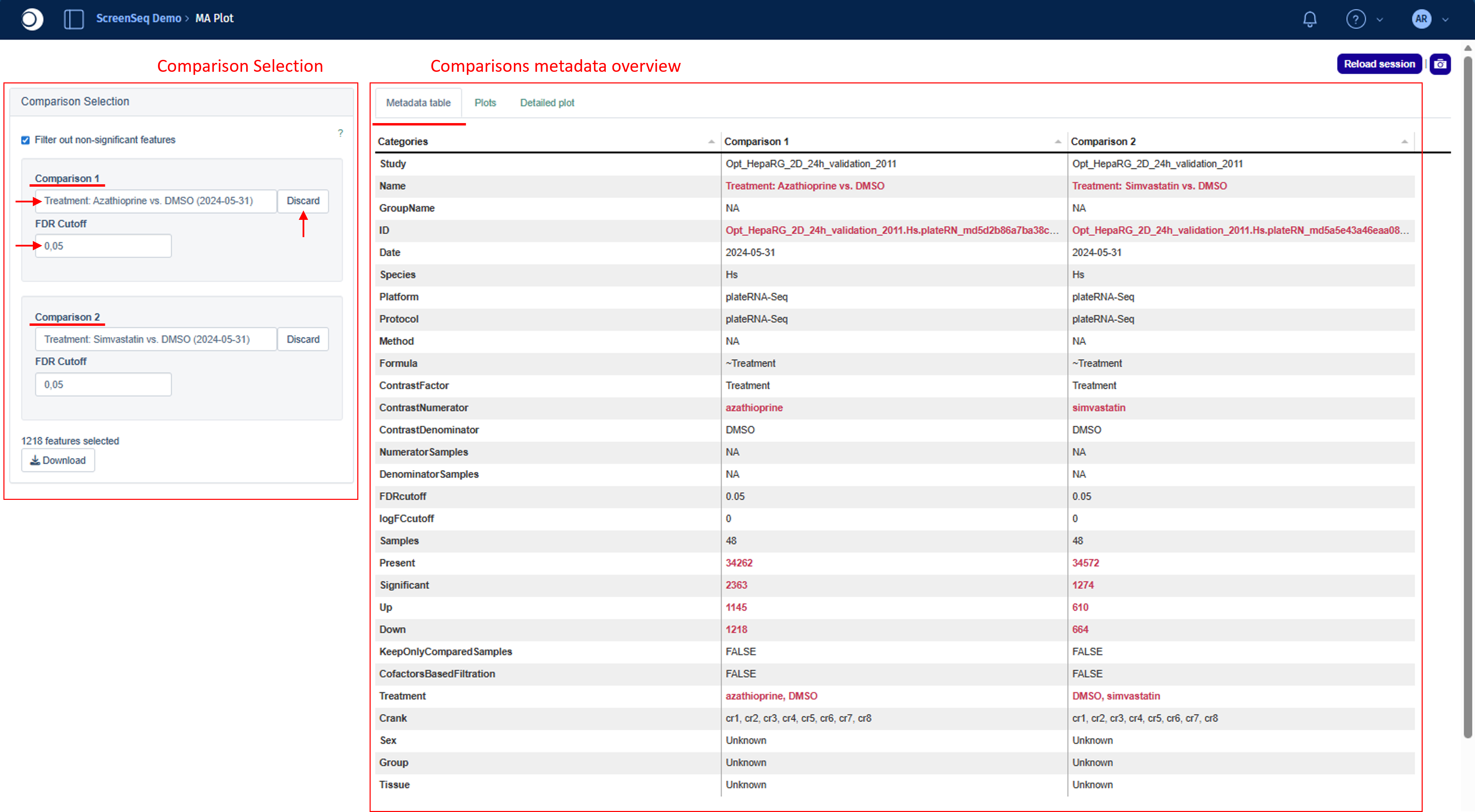
The Comparison Selection module provides two fields: Comparison 1 and Comparison 2. Clicking either field opens the Comparison Selector, where you can browse and select the desired comparison.
Note: A step‑by‑step guide to using the Comparison Selector is available under General Widgets.
If needed, you can remove the selected comparison at any time using the Discard button.
Adjusting FDR Thresholds
For each selected comparison, you can define an FDR threshold, which determines which genes are considered significantly differentially expressed. Each comparison has its own adjustable FDR field.
Based on this threshold:
- Non‑significant features are automatically filtered out
- Only features meeting the chosen significance criteria are displayed in the app
Note: Non‑significant features may still appear if they match a significant feature in the other selected comparison. Changing the FDR threshold for one MA plot can therefore alter the displayed features in both plots if previously hidden features become significant.
Turning Off Filtering
If you want to view all features (including non‑significant ones), you can disable filtering at any time by unticking the checkbox located at the top of the Comparison Selection module.
Metadata table view
Once you have selected the comparisons of interest, the Metadata table view in the results displays the combined metadata for both seected comparisons.
This table allows you to directly examine the similarities and differences between the selected comparisons in terms of their underlying recipe. It lists all selection criteria, the test formula, as well as the contrast numerator and denominator.
Any differences between the two comparison recipes are highlighted in bold red to make them easy to identify at a glance.
Why is comparing metadata important?
Comparing metadata ensures that differences observed in the MA Plot reflect true biological variation rather than inconsistencies in how the Comparisons were configured.
Please check the FAQ at the bottom for a more detailed explanation.
Interactive data exploration
Plots view
The Plots tab displays a combined feature table on the top and MA plot for each comparison. In addition, Plot options and Features to highlight are available below Comparison selection module on the right side.
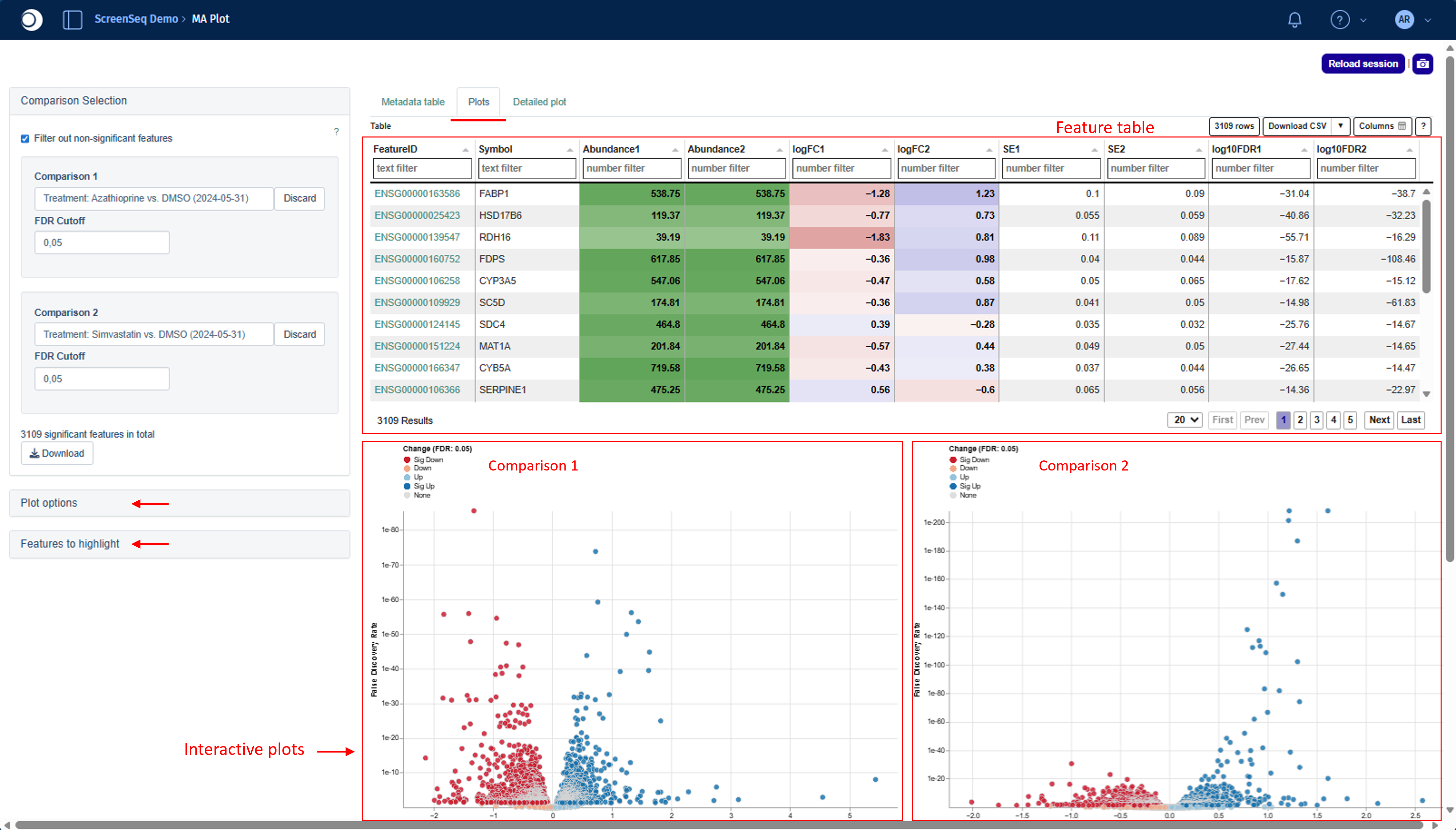
Feature table
The Feature table combines information from both selected comparisons. The columns shown depend on the underlying platform and data type.
The first two columns list the Feature IDs and Gene symbols. For proteomics or post‑translational modification (PTM) datasets — such as phosphoproteomics or ubiquitinomics — additional columns (e.g., UniProt IDs or PTM identifiers) are included.
For details on how features are matched across proteomics or peptidomics, please refer to the Algorithms section.
The remaining columns provide:
- Abundances of the denominator
- Log₂ fold changes (logFC)
- Standard error (SE)
- False discovery rate on log₁₀ scale (log10FDR)
Abundance values are shaded in light to dark green, where darker tones represent higher abundance. Similarly, logFC columns use red and blue shading:
- Red indicates down‑regulation (negative logFC)
- Blue indicates up‑regulation (positive logFC)
Additional table functionalities — such as filtering, sorting, and downloading — are described in the General Widgets section.
Interactive plots
Below the feature table, interactive plots, including Volcano/MA for each comparison separately, are displayed.
- Upper left - Volcano/MA plot for Comparison 1
- Upper right - Volcano/MA plot for Comparison 2
For every feature that is considered significantly differentially expressed in either comparison, a point is shown in both plots at the coordinates defined by that feature’s respective x‑ and y‑axis values.
All plots are fully interactive: you can zoom in/out, pan, select features, and view additional information on hover.
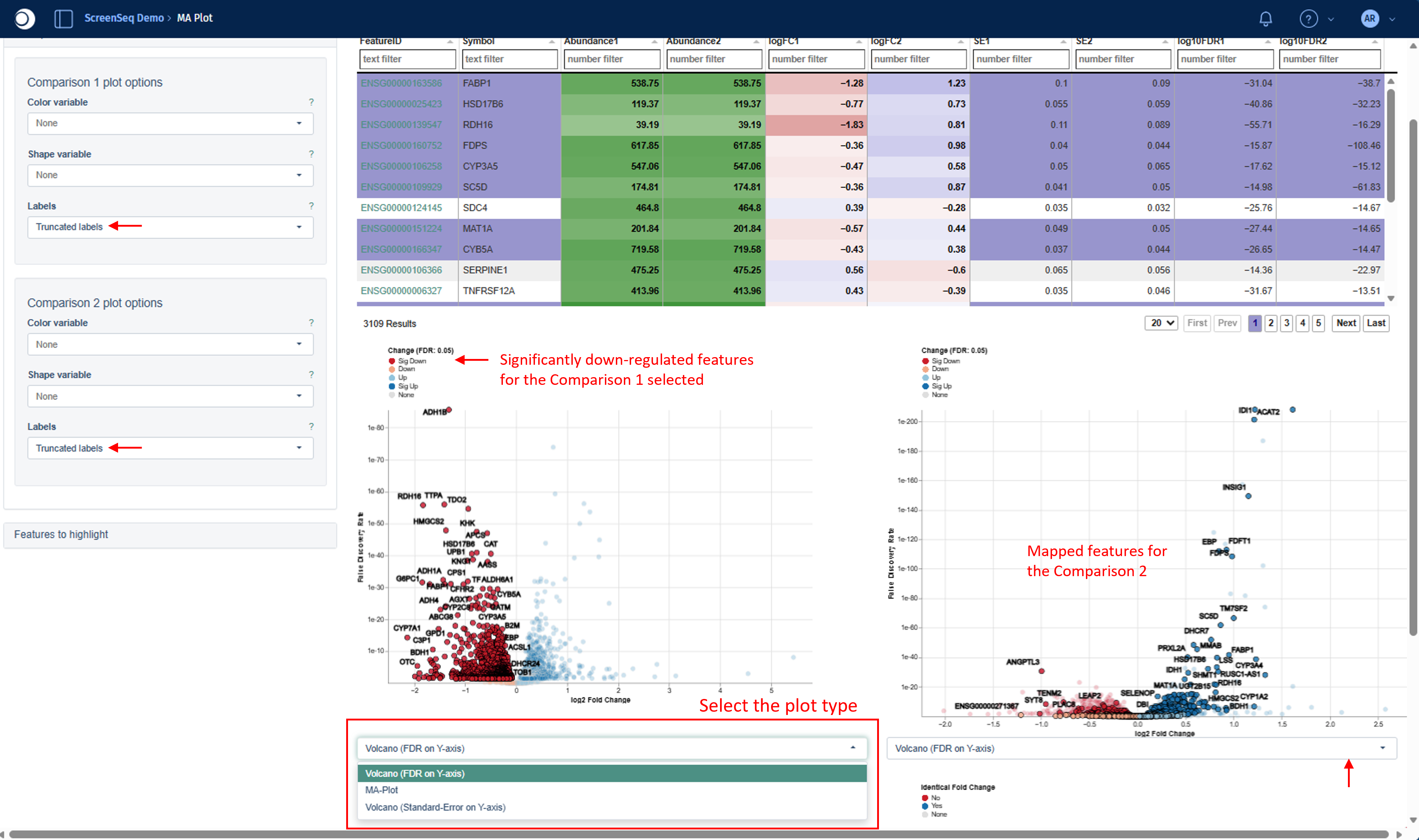
For each comparison, you can choose one of the following plot types using the drop‑down menu located directly below the plots:
- Volcano (logFC on x-axis, FDR on y-axis)
- Volcano (logFC on x-axis, Standard-Error on y-axis)
- MA Plot (Abundance on x-axis, logFC on y-axis)
On two Volcano/MA Plots, colors reflect the direction and statistical significance of regulation according to the applied FDR threshold:
- Dark red - significantly down-regulated
- Orange/Light red - down-regulated, but not significant
- Light blue - up-regulated, but not significant
- Dark blue - significantly up-regulated
- Grey - non-significant and unregulated
Below the individual Volcano/MA plots, two additional combined plots that visualize both Comparisons together in a single view are displayed:
- Bottom left - log₂FC (Comparison 1 on x‑axis, Comparison 2 on y‑axis)
- Bottom right - Feature abundances (Comparison 1 on x‑axis, Comparison 2 on y‑axis)
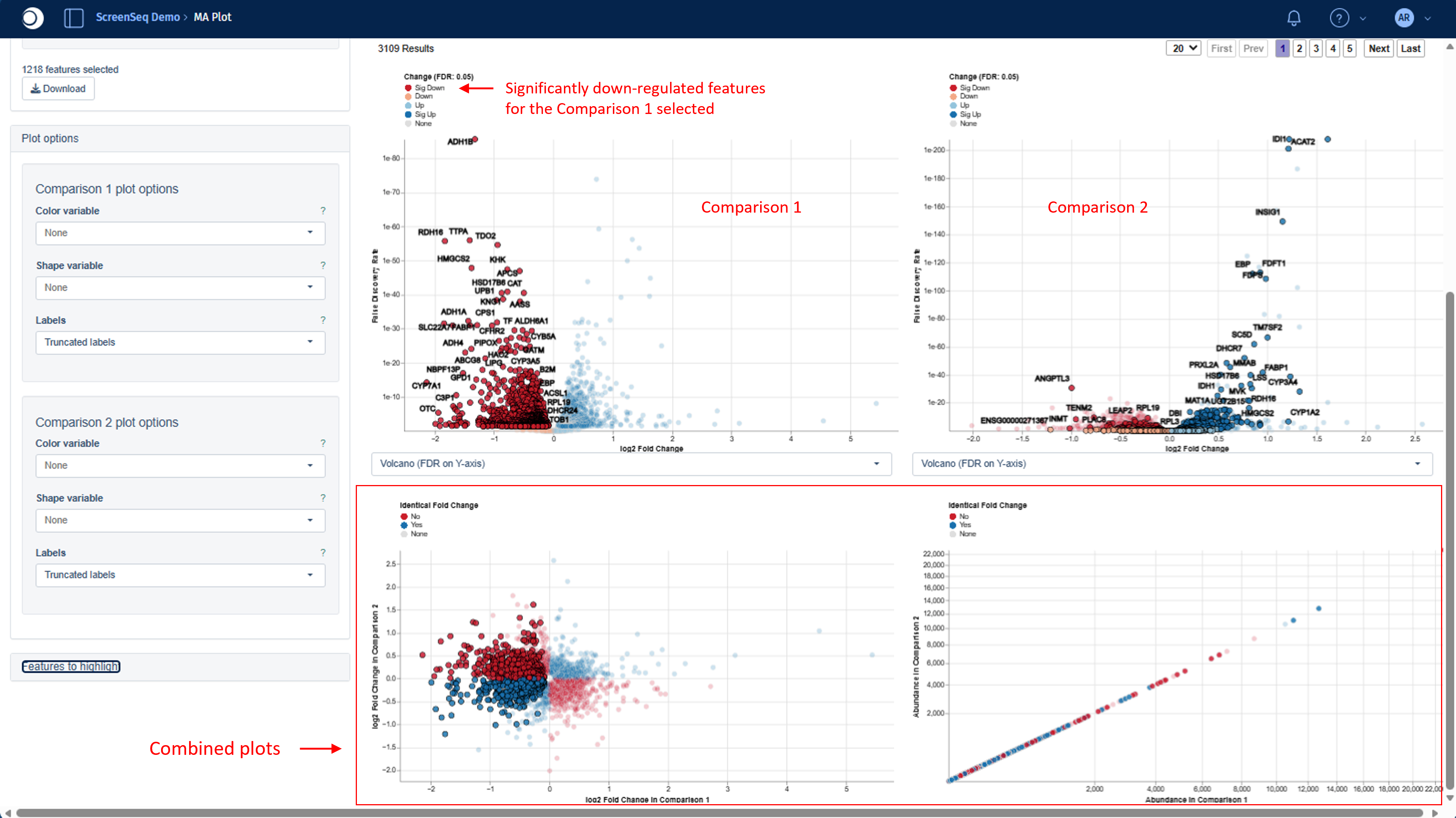
Colors indicate whether a feature shows the same direction of fold change in both Comparisons:
- Blue – Yes (both up‑regulated or both down‑regulated)
- Red – No (opposite directions; e.g., up in one, down in the other)
Feature selection
The app provides fully interactive tools for selecting and deselecting genes. Any feature selected in one view is automatically highlighted in all others — MA/Volcano plots, combined plots, and the Feature table — and the same applies in reverse.
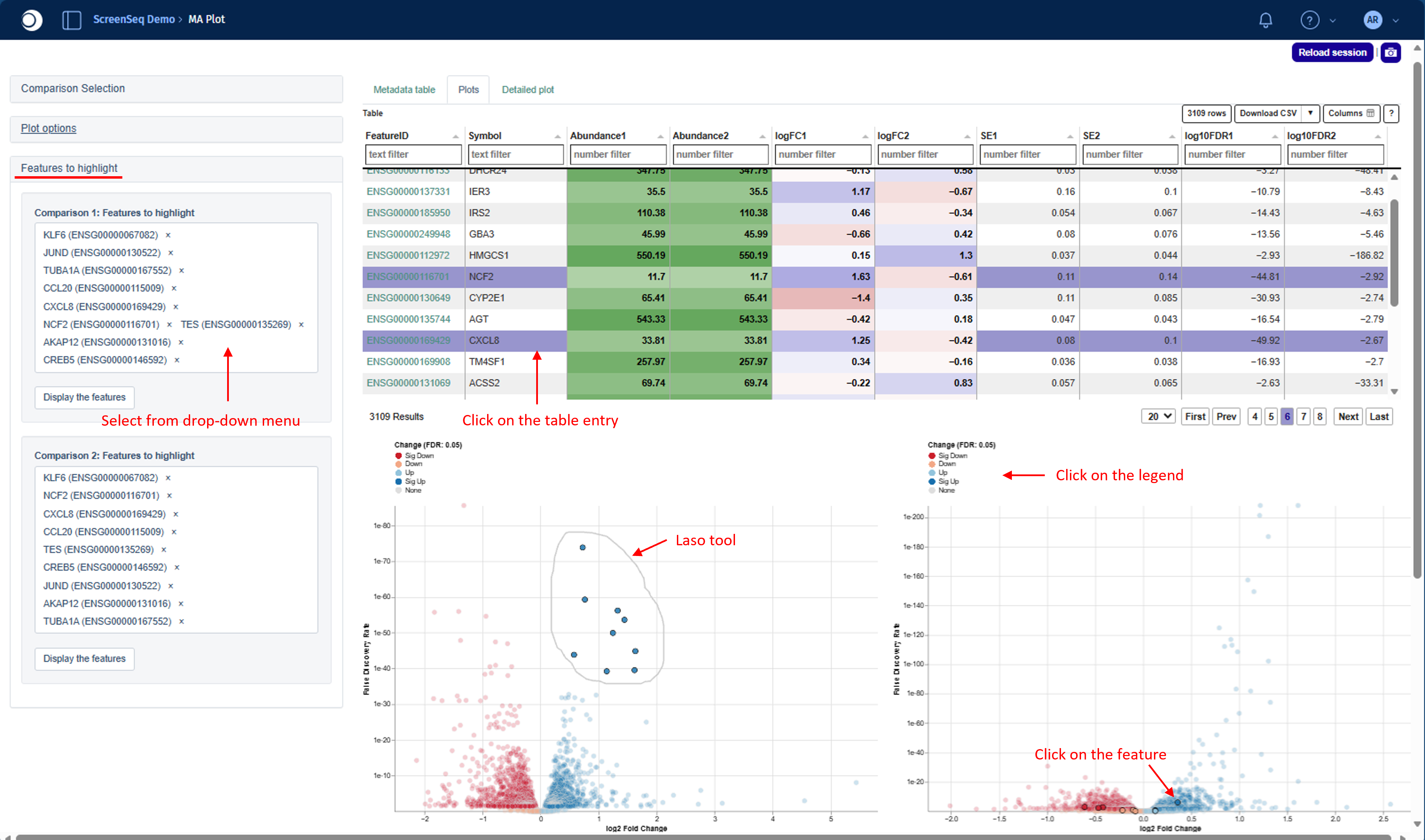
You can select features by:
- Clicking on one or more rows in the Feature table
- Clicking directly on feature points in any of the plots
- Clicking on values in the plot legend to highlight groups of features
- Using the laso tool
To activate laso tool please hold the right and circle the area on the plot - Selecting features from the drop-down menu in the Features to highlight module
Keyboard shortcuts support multi‑selection:
- CTRL – select multiple individual features
- SHIFT – select a continuous range
Selections are global across the app, meaning you can select features across multiple widgets simultaneously and see updates everywhere.
For example, you can select all significantly up‑regulated genes in one plot and combine them with those that are significantly up‑regulated in the other plot.
The Features to highlight module allows you to quickly select and display features of interest using a drop‑down menu. If features are selected through the table or directly from the plots, this field is automatically populated with the corresponding Feature IDs.
Plot Customization & Feature Annotation
The Plot options module lets you adjust point color and shape (depending on data type) and annotate selected features directly on the plot.
Color and shape settings are relevant when working with proteomics or PTM data, where these visual cues can reflect imputation or mapping status.
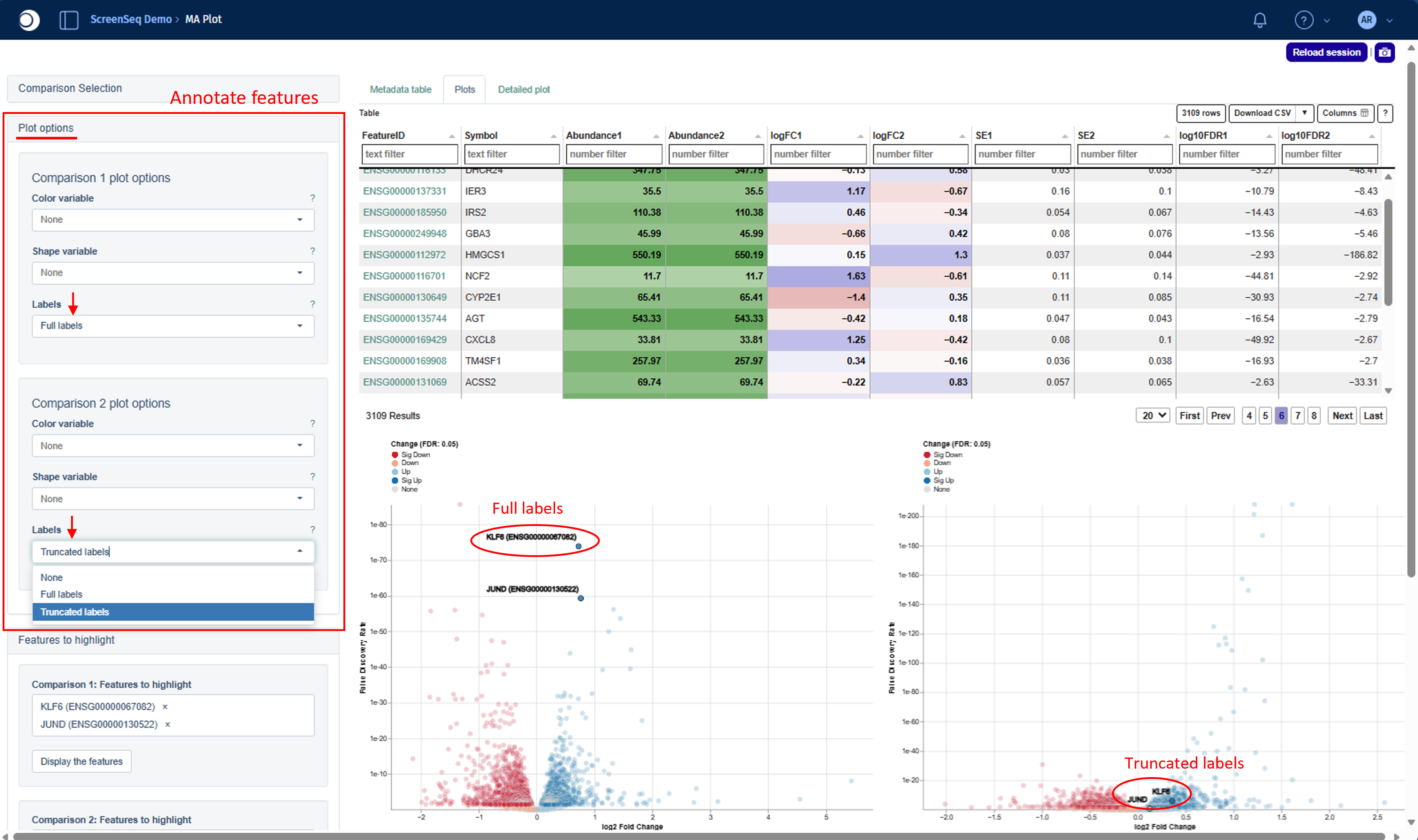
You can annotate selected features using:
- Full labels – EnsemblID or UniProtID + Gene Symbol
- Truncated labels – Gene Symbol only
Note: Settings in Plot options must be applied separately for each comparison plot.
A short app‑tour video at the top of the page provides a demonstration of all key features and interactions within the Plots tab.
Detailed plot view
The Detailed plot tab provides an in‑depth view of the abundance distribution across the contrast factor used in the selected Comparison.
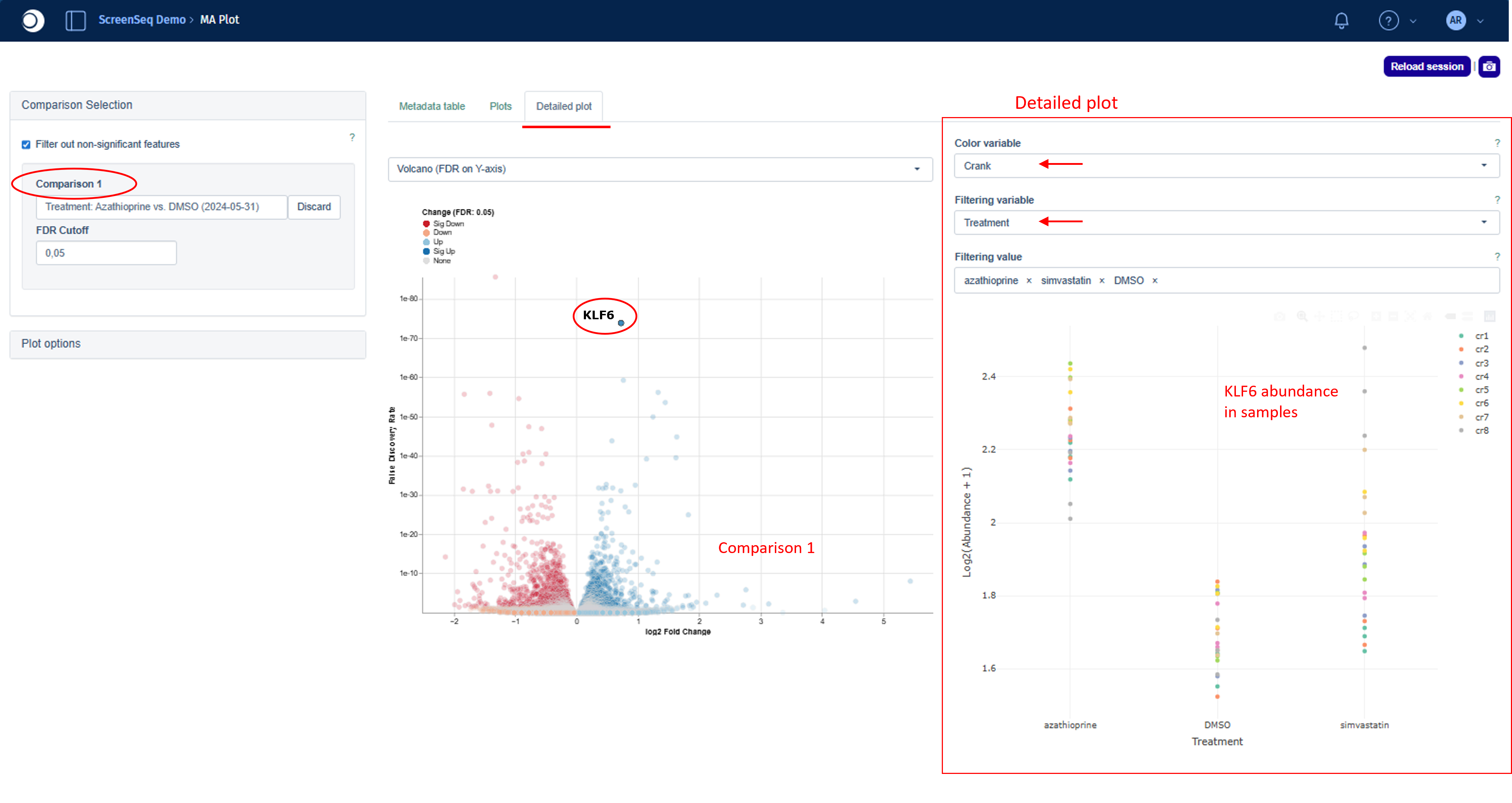
On the left side, the MA/Volcano plot for Comparison 1 is shown. When you select a single feature — either by clicking on it in the plot or by choosing it from the drop‑down menu in Plot options — a corresponding detailed abundance plot appears on the right.
For the selected feature, the plot displays abundance values in log-scale across the categories of the contrast factor. This helps you quickly understand what drives the observed fold change in the Comparison.
The plot can be customized further by adjusting colors or filtering samples. To filter out specific samples:
- Select a Filtering value (e.g. Sex)
- Choose one or more Filtering values to include (e.g. Female)
Filtering adjusts the detailed plot to show only the selected subset of samples.
Please note that the Detailed plot is not available for comparisons computed with the scExplorer app and custom comparisons generated outside of PanHunter.
Download features
Finally, a list of selected genes can be downloaded using the Download button available in the Comparison Selection module. The downloaded file reflects the exact set of features selected across the plots and the Feature table.